Volume 12 - Year 2025 - Pages 98-106
DOI: 10.11159/jbeb.2025.012
Biophysical Mapping of β-Lactamase–AuNP Conjugates: Mechanisms and Metrics for High-Resolution AMR Detection
Adwaita S R Nair1,2,3, Kunal Dhankhar1,4, Deepika Singh1,3, Arup Samanta2,3, Saugata Hazra*1,2
1Department of Biosciences and Bioengineering, Indian Institute of Technology Roorkee
Roorkee, Uttarakhand 247667, India
adwaita_n@nt.iitr.ac.in
2 Centre for Nanotechnology, Indian Institute of Technology Roorkee
Roorkee, Uttarakhand 247667, India
kunal@pe.iitr.ac.in
3 Department of Physics, Indian Institute of Technology Roorkee
Roorkee, Uttarakhand 247667, India
arup.samantha@ph.iitr.ac.in
4 Department of Polymer and Processing Engineering, Indian Institute of Technology Roorkee
Saharanpur, Uttarakhand 247667, India
saugata.iitk@gmail.com
*Corresponding Author: Dr. Saugata Hazra, Associate Professor, Department of Bioscience and Bioengineering, Indian Institute of Technology-Roorkee, Roorkee, Uttarakhand, India; Telephone no.: +918171064462; Email: saugata.iitk@gmail.com; saugata.hazra@bt.iitr.ac.in. ORCID ID: 0000-0002-3074-1534
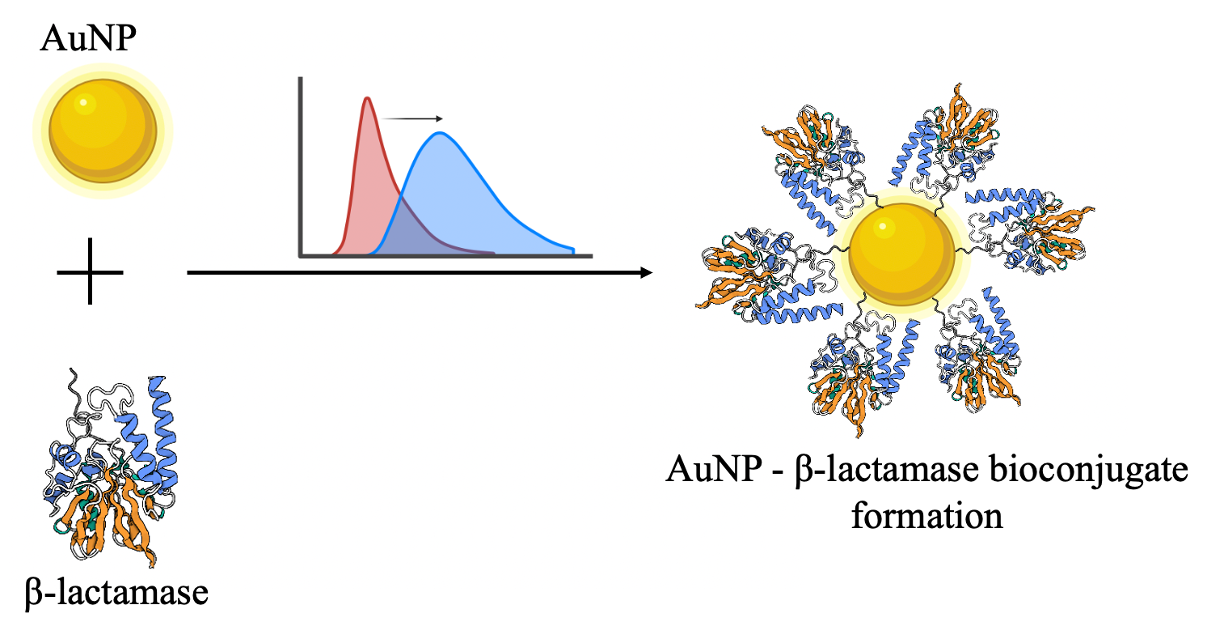
Abstract - With the increasing mortality attributed to antibiotic resistance, the development of a rapid and early detection biosensor for beta-lactamase enzyme identification has become imperative. The exceptional optical and electromagnetic characteristics of gold nanoparticles have rendered them as a prime candidate for biosensing applications. This research seeks to establish a foundation for the nanosensor development by investigating intricate interaction between gold nanoparticles and the beta-lactamase enzyme, focusing on functional and conformational dynamics following conjugation. UV-visible spectroscopy has been employed to examine the stability of bioconjugates and influence of pH on their conformational state by observing changes in the localized surface resonance plasmon band (LSRP). The minor red shift of the LSPR peak following beta-lactamase conjugation confirms protein conjugation and indicates the absence of gold nanoparticle aggregation due to the protein. The fluorescence quenching of tryptophan residues in beta-lactamase, in the presence of gold nanoparticles, is utilized to ascertain the binding of protein onto the surface. Circular dichroism spectroscopy further provided insight into the structural integrity of bioconjugate. Intact α-helix and beta-sheet peak in CD spectra confirmed that the interaction of gold nanoparticle surface did not cause unfolding or denaturation of protein. These complementary biophysical measurements, when combined, create a stable, non-denaturing conjugation system that produces strong optical signatures while keeping the structure of β-lactamase intact. This research provides essential insights into the interaction between beta-lactamase and gold nanoparticles, facilitating the advancement of sophisticated nanosensors that may be utilized in early pathogen detection and the monitoring of antimicrobial resistance.
Keywords: Gold nanoparticle, Beta-lactamase, Antibiotic resistance, Bioconjugate, FTIR
© Copyright 2025 Authors This is an Open Access article published under the Creative Commons Attribution License terms. Unrestricted use, distribution, and reproduction in any medium are permitted, provided the original work is properly cited.
Date Received: 2025-01-29
Date Revised: 2025-09-26
Date Accepted: 2025-10-15
Date Published: 2025-11-26
1. Introduction
Worldwide deaths due to the inability to treat diseases with current known antibiotics has brought inspection of antibiotic resistance to the limelight of research. According to a survey, deaths due to antimicrobial resistance have crossed 1.2 million as of 2019, thus bypassing the death counts due to HIV or malaria [1]. Multiple studies direct towards inefficiency of current treatments for more than half of the clinical patients. Rapid dissemination of antibiotic-resistant bacteria by horizontal gene transfer specifically carbapenemase resistant ESKAPE pathogen has raised concerns among all International and National health agencies, policymakers, and researchers. Over the years bacteria have adapted multiple methods to combat antibiotics such as pump efflux, reduction of drug permeation across the bacterial membrane, alteration of target sites and beta-lactamase expression. beta-lactam drugs being a major arsenal against antimicrobial resistance plays a major role in the development of antibiotic resistance in bacteria. [2], [3]. Hydrolysis of beta-lactam drug is one of the most concerning mechanisms adapted by bacteria to overcome the effect of antibiotics. Beta-lactamase genes are plasmid-encoded, thus rapidly escalating their presence in all the corners of the world. Their ability to cleave all classes of beta-lactam drugs including carbapenem, which is usually the last resort, and the absence of clinically accepted inhibitors makes the understanding its study need of the hour[4] The concerning rate of beta-lactamase spreading across the world directly points at the urgent need for deep understanding and remedy for the same. Thus, thorough literature search and mechanistic study of emergence, activity, spreading and remedies for more beta-lactamase should be foremost and noteworthy. Colloidal gold nanoparticles (AuNPs) have attracted considerable interest in recent years due to their remarkable characteristics, including size and shape-dependent optoelectronic properties, an extensive surface-to-volume ratio, minimal toxicity, and good biocompatibility. Researchers globally have advanced in regulating its dimensions, morphology, and functionalization, alongside various synthesis techniques. Distinct characteristics, including surface plasmon resonance, size-dependent absorption color, and fluorescence quenching capability, significantly enhance their extensive utility in diagnostics, treatments, and imaging.[5] The simplicity of functionalization and biocompatibility render gold nanoparticles exceptionally advantageous for interaction with biomolecules, including enzymes. Research on bioconjugate nanoparticles has garnered significant attention due to their potential uses in luminescence tagging, imaging, medical diagnostics, multiplexing, and biosensors. The binding of protein to the nanoparticle surface may significantly affect its morphology and functionality. Park [5] examined the significance of understanding the conformational changes and unfolding mechanisms of proteins for the enhancement of nanoparticles' biomedical applications. Beta-lactamase protein is a major concern and has been the focus of research, but not much has been done to study how it interacts with nanoparticles. This limits the usage of nanoparticle for sensing and therapeutics of antibacterial resistance associated beta-lactamase protein. While the interaction of proteins such as lysozyme, BSA, etc. have been explored, the possibility of interaction with beta-lactamase protein has not been yet explored. Considering, SME-1 to be a fastest carbapenemase reported in the literature, it has been chosen as the representative candidate for beta-lactamase enzyme. In this study, we aim to ascertain if beta-lactamase protein may adhere to the surface of gold nanoparticles. We investigate the conformational alterations experienced by the protein upon binding and whether this interaction may induce any denaturation of the protein.
2.Material and methods
2.1 Chemicals
Hydrogen tetrachloroaurate (III) trihydrate (HAuCl₄·3H₂O, ≥99.9% trace metals basis) and trisodium citrate dihydrate (Na₃C₆H₅O₇·2H₂O, analytical grade) were purchased from Sigma Aldrich. All chemicals were used without further purification. Ultrapure deionized water (resistivity ≥18.2 MΩ·cm) was used in all experiments.
2.2 Gold nanoparticle synthesis and characterisation
Gold nanoparticles were synthesized by a method initially introduced by Turkevich and further modified by Frens [7], [8]. In this method, aqueous solution of HAuCl₄ (1mM) was heated to 95 °C under constant stirring. After 10 min, the gold solution was reduced by the rapid addition of Na₃C₆H₅O₇ (38.8mM). This mixture was further heated under constant stirring. After the confirmation of gold nanoparticle synthesis by change of color to ruby red from yellow, it was further heated for 10 min and cooled to room temperature. The synthesised gold nanoparticle was refrigerated at 4 °C in glass vials until further requirement. Agilent Cary 60 UV-Vis spectroscopy was used to evaluate the absorption spectrum of as-synthesised gold nanoparticle.[9] The surface morphology and size synthesized gold nanoparticles were assessed using a field emission scanning electron microscope (FESEM, Apreo S LoVac, Thermo Fischer Scientific) and transmission electron microscopy (TEM TALOS F200-X).[10] These images were then evaluated using ImageJ software for their size.[11] The hydrodynamic diameter of gold nanoparticle was determined using Malvern zetasizer.
2.3 Cloning and purification of SME-1 beta-lactamase
The SME-1 gene from Serratia marcescens was cloned and expressed using the pET-28a(+) vector. The gene sequence for SME-1 was initially retrieved from the UniProt database, and the cloning was planned as shown in the Fig. 1a. The cloning was done using restriction enzymes Ncol and Xhol. The cloned plasmid expressing the N-terminal 6 histidine-tagged SME-1 protein was transformed into competent Escherichia coli BL21 (DE3 cells). Their overexpression was optimized using six different IPTG concentrations (0.1 mM, 0.5 mM, 1 mM) and temperatures (16°C, 18°C, and 22°C). Protein expression was performed at 0.5mM IPTG concentration after achieving an absorbance of 0.6 at 600 nm (OD600). After 16 hours of incubation at 18°C, cells were harvested and resuspended in a pH 7.5 buffer containing 55 mM Tris-HCl and 250 mM NaCl. To prevent proteolytic activity, a protease inhibitor was added to the resuspended solution. The resuspended solution was lysed using a sonicator, which were then centrifuged. The supernatant was filtered through a 0.45-μm syringe filter and put onto a Ni-NTA agarose column for affinity chromatography. The protein was eluted in 55 mM Tris HCl (pH 7.5) with 250 mM NaCl in steps of 10 mM to 250 mM imidazole. Furthermore, the 250 mM imidazole-eluted protein was concentrated using a 50 kDa concentrator. The concentrated protein was placed onto a Superdex 75 pg column (120 mL capacity) for size exclusion chromatography utilizing the AKTA start purification system. The pure protein was further concentrated, and its purity was determined by SDS-PAGE. 3D structure of the protein was visualised by using Pymol 3.1. Secondary structure analysis using circular dichroism (CD) spectroscopy were carried out in a Jasco J-1500 spectrometer with a Peltier-effect temperature controller. Quartz cells with a 0.1 cm path length were used.
2.3 Bioconjugate synthesis and characterisation
Histidine-tagged SME-E166A was incubated with citrate stabilized gold nanoparticles to allow their interaction.[12] To optimize their conjugate condition, they were incubated in varying molar ratio of gold nanoparticle to protein. This mixture was incubated for 1 hour under mild stirring to allow coordination between imidazole group and nanoparticle surface. Malvern zetasizer was used to evaluate the surface charge changes on gold nanoparticle with protein interaction. All the measurements were done at 25 °C. The interaction of protein and gold nanoparticle was further characterised by using protein’s intrinsic tryptophan fluorescence property. The bioconjugate samples were studied using Horibo fluorescence spectroscopy. The samples were excited at 295 nm and their emission was recorded for 310-550 nm. UV-Vis spectroscopy was used to understand the shift in SPR as discussed before. The change in secondary structures in protein after nanoparticle conjugation was evaluated using circular dichroism spectroscopy as well.
3. Result and discussion
3.1 Physicochemical Characterization of Synthesized Gold Nanoparticles
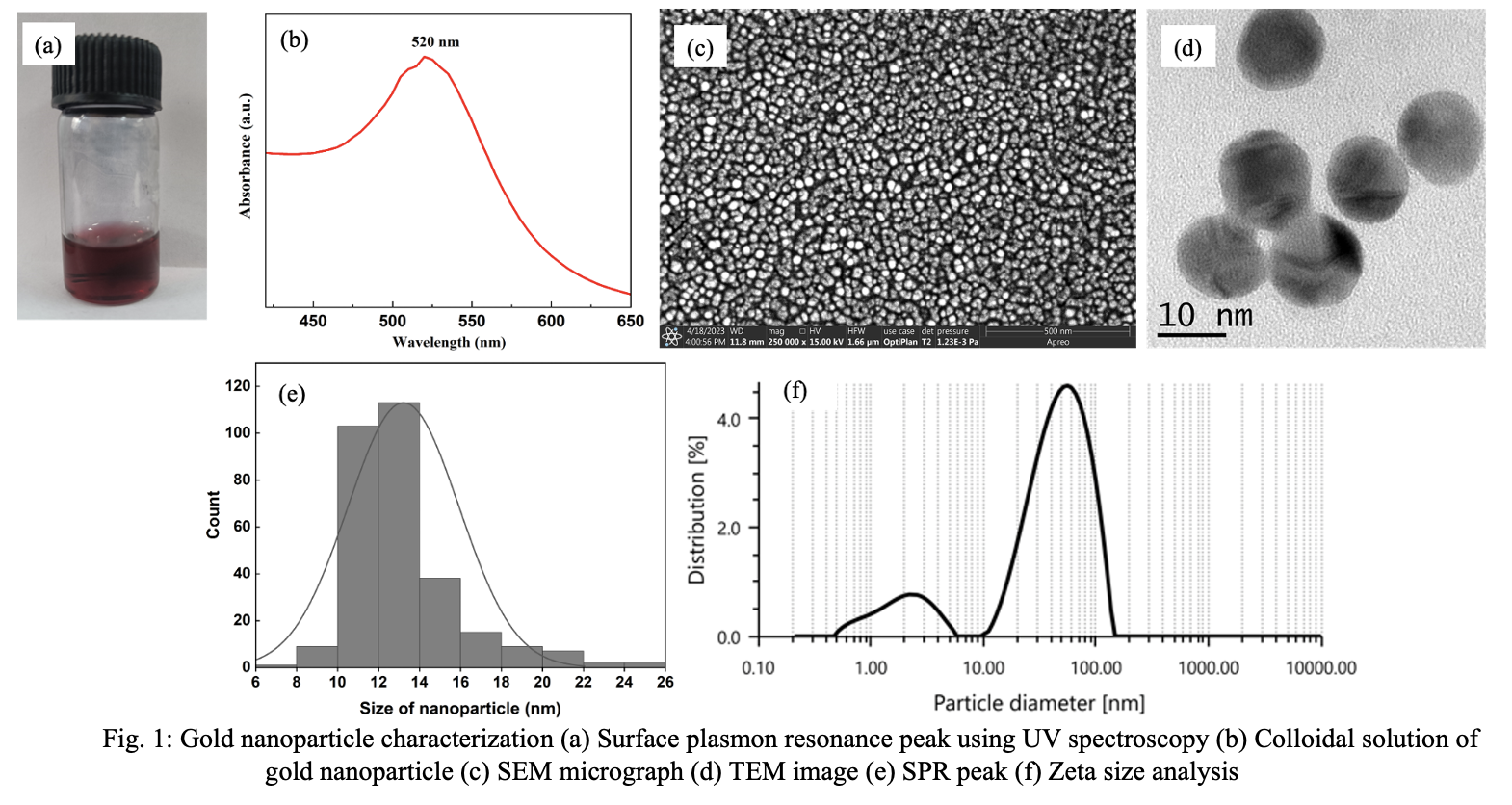
Fig. 1 illustrates the physicochemical characterization of the synthesized gold nanoparticles, employing various techniques to assess their properties. The color of the colloidal solution showed the characteristic wine-red color, which suggested synthesis of non-aggregated gold nanoparticle (Fig. 1a). UV-Vis absorption spectrum of gold nanoparticle displayed a distinct absorption peak at 520 nm (Fig. 1b). This agrees with previous literature, wherein gold nanoparticle of 10-20 nm size shows distinct peak at 518-524 nm.[13] This intense absorption peak could be attributed to the electronic oscillation i.e. surface plasmon resonance (SPR). The symmetric SPR peak might be indicative of monodisperse distribution of gold nanoparticle in the solution. Fig. 1c and Fig. 1d presents morphological analysis of gold nanoparticles synthesized via the Turkevich method. The SEM micrograph reveals a uniform dispersion of gold nanoparticles between 12-16 nm size with minimal agglomeration. Additionally, the size of the spherical gold nanoparticles was averaged to be 13 nm. (Fig. 1e) The relatively narrow size distribution highlights the effectiveness of citric acid as reducing agent and stabilizing agent. Fig. 1f depicts the
particle size distribution curve obtained through dynamic light scattering (DLS), indicating moderate polydispersity with a PDI of 0.272 and a hydrodynamic size of 38.3 nm (Fig. 1f). The higher hydrodynamic size is contributed by metal core, citrated capping and solvent shell. The presence of small peak at smaller size, indicate sub-population of smaller particles. The surface charge measurement, as assessed by DLS, was -32.6 mV, reflecting their stability in colloidal state.
3.2 Characterization of SME beta-Lactamase
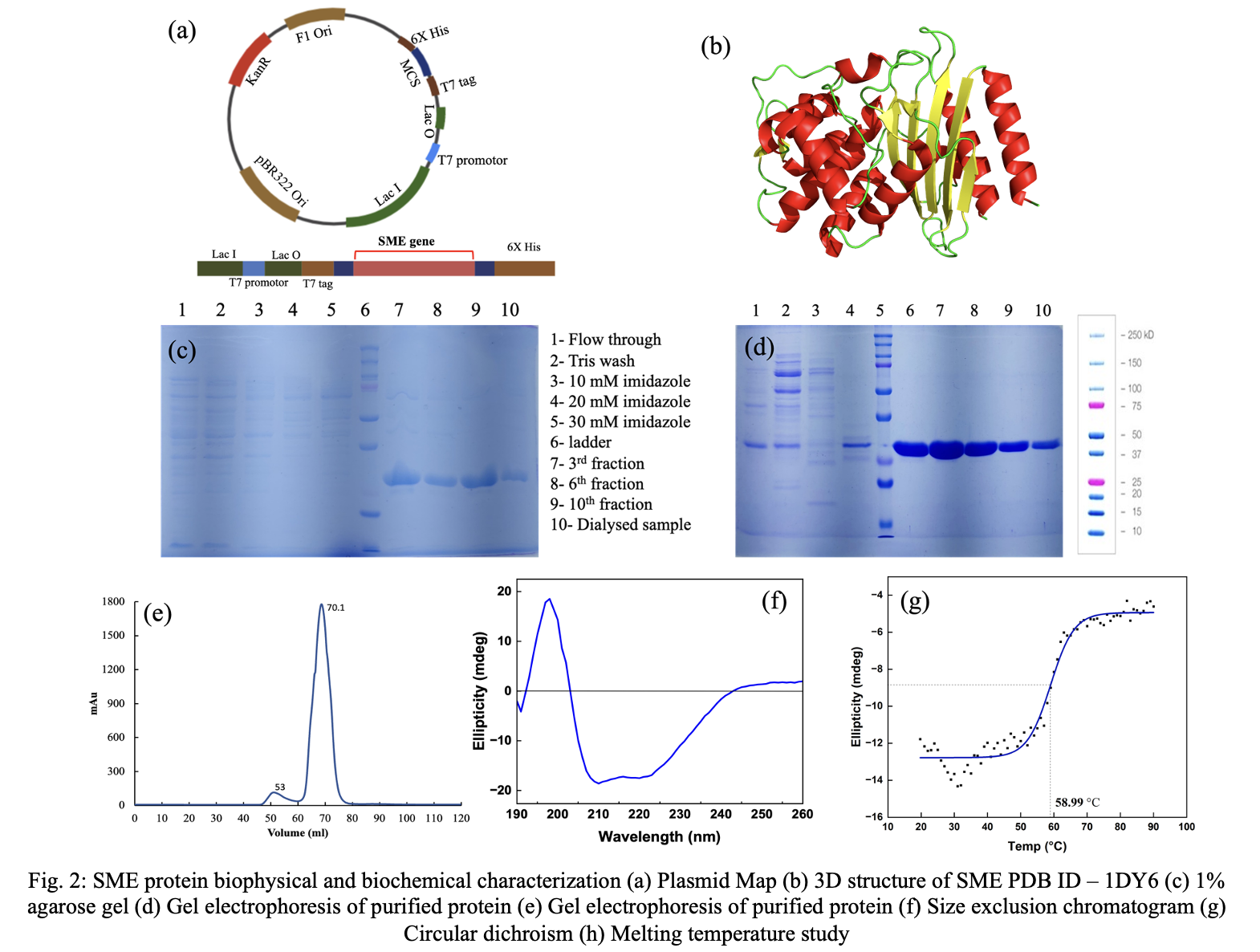
SME, a common beta-lactamase from Serratia marcescens, was chosen because it has classical carbapenemase activity and can showcase a range of catalytic capabilities. Using the NcoI and XhoI restriction enzymes, the SME gene was cloned into the pET-28a (+) vector (Fig. 2a). The vector map shows elements such as origin of replication, multiple cloning site, antibiotic resistance marker among many other. 3D structure of SME-1 protein was investigated by extracting from RCSB PDB (1DY6). The 3D structure of SME-1 enzyme revealed characteristic beta-lactamase fold αββα, wherein β-sheets are flanked by α-helices as shown in (Fig. 2b). SME protein was purified by two-step purification process, Ni-NTA agarose followed by size exclusion chromatography.[14] The purified proteins were analyzed using gel electrophoresis, which revealed protein bands at approximately 28 kDa. (Fig. 2c and Fig. 2d) The size exclusion chromatogram displayed a highly intense singular peak at 65 mL corresponding to SME protein, which was collected for subsequent characterization. (Fig. 2e) The presence of a single peak
indicated presence of protein in monomeric and homogenous state. The protein's secondary structure was assessed by circular dichroism spectroscopy. (Fig. 2f) The CD spectrum indicates that the SME beta-lactamase possesses a distinct secondary structure, predominantly characterized by a combination of alpha-helices and beta-sheets. These features are comparable to the components observed in model structure. Furthermore, the melting temperature (Tₘ) of SME was determined using CD spectroscopy to evaluate its thermal stability.[15] Figure 2g shows that the Tₘ of SME is about 55°C. This is the temperature at which fifty percent of the protein undergoes unfolding. The sigmoidal unfolding profile shows that the native state is well-folded and goes through a clear transition instead of slowly coming together. This makes the folding of SME-1 even more likely to be correct.
3.3 Evaluation of Bioconjugate Under Variable Conditions
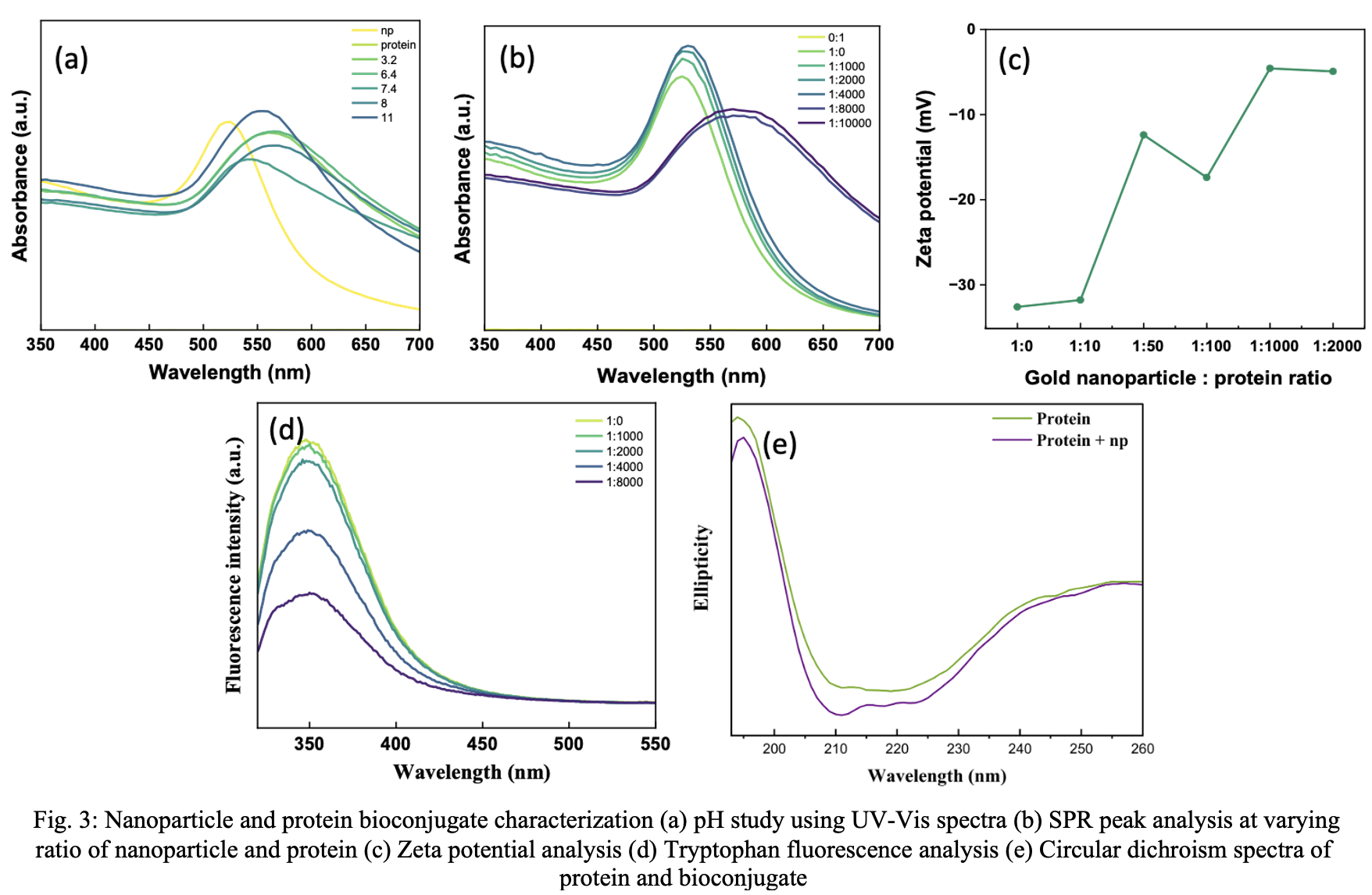
The SPR peak spectra were utilized to evaluate the in-situ conditions of bioconjugate formation at each pH level (3, 6, 7, 8, 11) examined in this investigation. A shift in the SPR peak signifies alterations
in the chemical environment surrounding gold nanoparticles, hence allowing its assessment to correspond with protein interactions. Fig. 3a illustrates that citric acid-capped gold nanoparticles have a surface plasmon resonance peak at 540 nm, while the protein only had baseline absorption in UV region. The presence of protein alters both the peak's broadness and its position.[16] The red shift, indicative of protein desorption or aggregation, suggests that an acidic media is unsuitable for bioconjugation. At elevated pH levels, the SPR peak exhibits a minor red shift accompanied by considerable broadening. Consequently, a pH range of 7-8 may be deemed optimal for the enhanced stability of the bioconjugate. Further using the same UV-Vis spectral approach, the optimum np to protein ratio was studied. Up until 1:1000 ratio of np to protein, the SPR peak doesn’t show significant shift. (Fig. 3b) But further increase shows shifting and broadening of SPR peak indicating aggregation of gold nanoparticle. This suggest that bioconjugates are not stable at higher concentration ratios.
3.4 Fluorescence Quenching Analysis
Quenching of tryptophan fluorescence in presence of gold nanoparticle was investigated to better understand the interaction and quenching mechanism of bioconjugate interaction.[17] While the maximum emission wavelength (about 350nm) was nearly same in all cases (fig. 3c), fluorescence intensity decreased as with increasing GNP concentration which could be ascribed to gold nanoparticle quenching. The degree to which protein was conjugated with gold could be used to directly correlate to the degree at which fluorescence intensity was reduced.[18] This confirms the interaction of nanoparticle to the protein residues.
3.5 Zeta potential analysis
The zeta potential was employed to monitor the change in surface charge after protein conjugation to the nanoparticle surface as it can provide insights into the stability of bioconjugate.[19] At 1:0 ratio, the gold nanoparticle seems to have a surface charge of approximately -35 mV, which could be attributed to the citrate group used as capping agent. (fig. 5d) With increasing protein concentration, zeta potential increases towards positive charge. This indicates interaction of positively charged residues with the nanoparticle surface. At higher concentration ratio (of above 1:1000) there is no further change in the surface charge as no further absorption of protein occurs. After 1:100 nanoparticle to protein ration, the zeta potential increase to more positive value >-10 mV, suggesting instability. Thus, zeta potential study corroborates with UV-Vis study, indicating instability at higher ratio.
3.6 Structural Integrity Assessment
Circular spectroscopy was used to assess whether beta-lactamase was able to retain its native structure after conjugation. As discussed earlier also, spectrum in fig. 5e illustrates the inherent structure of the protein. The peak at 195–200 nm shows the existence of beta-sheet or random coil structures, whereas the dip at 210–220 nm signifies α-helical content. The lack of a significant shift in the ellipticity peak and a decrease in intensity indicate the presence of intact beta-lactamase protein following nanoparticle conjugation. The spectral study indicate that the protein doesn’t undergo much conformational alteration upon bioconjugation.
4. Conclusion
The increasing prevalence of antibiotic resistance resulting in fatalities has brought beta-lactamase to the forefront. These beta-lactam hydrolyzing proteins have been in the limelight of research, since the recognition of AMR as global threat by UNESCO. The increasing prevalence of newly found beta-lactamase mutations has once more elicited worries among researchers and health officials. These new mutations pose a threat for the development of suitable countermeasures. With the advent of nanotechnology, many nanoparticles are utilized for the detection and theranostics of diseases. Among these gold nanoparticles stand out owing to its remarkable properties such as surface plasmon resonance, size-based properties and ease of synthesis. Their long history in biosensing gives them a strong methodological base. Their size-dependent physics and simple surface chemistry make them easy to customize. Their bright, tuneable surface-plasmon resonance turns tiny molecular events into clear optical signatures. In this study, we have sought to comprehend the interaction between gold nanoparticles and the beta-lactamase protein, delving into the molecular mechanism underpinning their conjugation. This toolkit was made to answer three main questions: does the enzyme stick to the particle, does the particle report that binding accurately, and does binding keep the enzyme's native fold?
Gold nanoparticles, 13 nm in size were assessed and appear to interact with beta-lactamase protein at a pH of 6-7, with no alteration to its secondary structure. Fluorescence spectroscopy shows that intrinsic tryptophan emission, which is a close, distance-dependent indicator that the protein is getting close to and sticking to the nanoparticle surface, is clearly quenched when incubated with AuNPs. Surface-charge measurements back up this close relationship by showing that there is a stable interfacial complex instead of a short interaction. Importantly, circular dichroism spectra show that canonical α-helical and β-sheet signals are still there after conjugation. This strongly suggests that denaturation or unfolding did not happen in our case. While this study could serve as a foundation for the advancement of sensors utilizing gold nanoparticles, further kinetic study needs to be evaluated to understand the functional changes upon bioconjugate formation. In in-vivo condition serum protein may also bind to gold nanoparticle surface and thus hinder the binding of beta-lactamase with gold nanoparticle. At the same time, salts may vary the pH and biological fluids may present different physiological environment, which may not be suitable for bioconjugate stability. Thus, since these experiments were performed in controlled environment, the extent at which bioconjugates are affected in presence of other protein, salts and biological fluids must be explored. We also need to investigate how the size, shape, and functionalization of AuNPs affect sensitivity and specificity, how to make surface chemistries work better for repeatable conjugation densities, and how to test stability in complex biological matrices. This research offers a critical instrument in the fight against antibiotic resistance by utilizing the distinctive properties of AuNPs to develop innovative solutions for the detection and monitoring of beta-lactamase activity.
Acknowledgements
Authors thankfully acknowledge Centre for Nanotechnology, IIT Roorkee and Indian Institute of Technology, Roorkee for the support and facilitating necessary infrastructure. We acknowledge the Board of Research in Nuclear Sciences (BRNS) for their support under Sanction Number 54/14/03/2023-BRNS/12162 and the Indian Council of Medical Research (ICMR) for their grant under ISRM Sanction Number 2023-0773. A.S.R.N acknowledges the Ministry of Education, Government of India for her doctoral fellowship through the Prime Minister's Research Fellowship (PMRF) scheme.
References
[1] T. Thompson, 'The staggering death toll of drug-resistant bacteria', Nature, Jan. 2022, doi: 10.1038/D41586-022-00228-X. Available: View Article
[2] S. Bhattacharya, V. Junghare, N. K. Pandey, D. Ghosh, H. Patra, and S. Hazra, 'An insight into the complete biophysical and biochemical characterization of novel class A beta-lactamase (Bla1) from Bacillus anthracis', Int J Biol Macromol, vol. 145, pp. 510-526, Feb. 2020, doi: 10.1016/J.IJBIOMAC.2019.12.136. Available: View Article
[3] S. Bhattacharya, A.K. Padhi, V. Junghare, N. Das, D. Ghosh, P. Roy, K. Y. J. Zhang and S. Hazra, 'Understanding the molecular interactions of inhibitors against Bla1 beta-lactamase towards unraveling the mechanism of antimicrobial resistance', Int J Biol Macromol, vol. 177, pp. 337-350, Apr. 2021, doi: 10.1016/J.IJBIOMAC.2021.02.069. Available: View Article
[4] K. C. A. Kumar, A. Nair, S. Sharma, D. Singh, S. Yadav, D. Bhimsaria, S Gupta and S. Hazra, 'Strategic design of a multi-tier database for class A $\beta$-lactamase BlaC variants of M. tuberculosis: advancing the fight against antibacterial resistance', The Journal of Antibiotics 2025, pp. 1-16, Aug. 2025, doi: 10.1038/s41429-025-00862-3. Available: View Article
[5] Y. C. Yeh, B. Creran, and V. M. Rotello, 'Gold Nanoparticles: Preparation, Properties, and Applications in Bionanotechnology', Nanoscale, vol. 4, no. 6, p. 1871, Mar. 2011, doi: 10.1039/C1NR11188D. Available: View Article
[6] S. J. Park, 'Protein-nanoparticle interaction: Corona formation and conformational changes in proteins on nanoparticles', Int J Nanomedicine, vol. 15, pp. 5783-5802, 2020, doi: 10.2147/IJN.S254808. Available: View Article
[7] J. Turkevich, P. C. Stevenson, and J. Hillier, 'A study of the nucleation and growth processes in the synthesis of colloidal gold', Discuss Faraday Soc, vol. 11, pp. 55-75, 1951, doi: 10.1039/DF9511100055. Available: View Article
[8] J. Kimling, M. Maier, B. Okenve, V. Kotaidis, H. Ballot, and A. Plech, 'Turkevich method for gold nanoparticle synthesis revisited', Journal of Physical Chemistry B, vol. 110, no. 32, pp. 15700-15707, Aug. 2006, doi: 10.1021/JP061667W/ASSET/IMAGES/LARGE/JP061667WF00009.JPEG. Available: View Article
[9] A. S. R. Nair, S. Devi, S. Mandal, U. K. Tripathi, D. Roy, and N. E. Prasad, 'Insights into enzymatic degradation of physically crosslinked hydrogels anchored by functionalized carbon nanofillers', New Journal of Chemistry, vol. 46, no. 6, pp. 2669-2677, Feb. 2022, doi: 10.1039/D1NJ04924K. Available: View Article
[10] A. Singh, V. C. Srivastava, and I. Janowska, 'Utilization of carbon-black industry waste to synthesize electrode material for supercapacitors', Energy Storage, vol. 6, no. 5, p. e677, Aug. 2024, doi: 10.1002/EST2.677. Available: View Article
[11] M. Dhiman, S. Ghosh, T. G. Singh, S. Chauhan, P. Roy, and D. Lahiri, 'Exploring the potential of an Aloe vera and honey extract loaded bi-layered nanofibrous scaffold of PCL-Col and PCL-SBMA mimicking the skin architecture for the treatment of diabetic wounds', J Mater Chem B, vol. 12, no. 40, pp. 10383-10408, Oct. 2024, doi: 10.1039/D4TB01469C. Available: View Article
[12] R. Peltomaa, F. Amaro-Torres, S. Carrasco, G. Orellana, E. Benito-Pena, and M. C. Moreno-Bondi, 'Homogeneous Quenching Immunoassay for Fumonisin B1 Based on Gold Nanoparticles and an Epitope-Mimicking Yellow Fluorescent Protein', ACS Nano, vol. 12, no. 11, pp. 11333-11342, Nov. 2018, doi: 10.1021/ACSNANO.8B06094/ASSET/IMAGES/MEDIUM/NN-2018-06094X_M001.GIF. Available: View Article
[13] W. Haiss, N. T. K. Thanh, J. Aveyard, and D. G. Fernig, 'Determination of size and concentration of gold nanoparticles from UV-Vis spectra', Anal Chem, vol. 79, no. 11, pp. 4215-4221, Jun. 2007, doi: 10.1021/AC0702084/SUPPL_FILE/AC0702084SI20070321_014144.PDF. Available: View Article
[14] S. Bhattacharya, V. Junghare, N. K. Pandey, S. Baidya, H. Agarwal, N.Das, A. Banerjee, D. Ghosh, P. Roy, H. K. Patra and S. Hazra 'Variations in the SDN Loop of Class A Beta-Lactamases: A Study of the Molecular Mechanism of BlaC (Mycobacterium tuberculosis) to Alter the Stability and Catalytic Activity Towards Antibiotic Resistance of MBIs', Front Microbiol, vol. 12, p. 710291, Oct. 2021, doi: 10.3389/FMICB.2021.710291/BIBTEX. Available: View Article
[15] S. Bhattacharya, A. K. Nautiyal, R. Bhattacharjee, A. K. Padhi, V. Junghare, M. Bhambri, D. Dasgupta, K. Y. J. Zhang, D. Ghosh and S. Hazra 'A comprehensive characterization of novel CYP-BM3 homolog (CYP-BA) from Bacillus aryabhattai', Enzyme Microb Technol, vol. 148, p. 109806, Aug. 2021, doi: 10.1016/J.ENZMICTEC.2021.109806. Available: View Article
[16] M. Iosin, V. Canpean, and S. Astilean, 'Spectroscopic studies on pH- and thermally induced conformational changes of Bovine Serum Albumin adsorbed onto gold nanoparticles', J Photochem Photobiol A Chem, vol. 217, no. 2-3, pp. 395-401, Jan. 2011, doi: 10.1016/J.JPHOTOCHEM.2010.11.012. Available: View Article
[17] A. Sharma, H. Rastogi, and K. Sundar Ghosh, 'Enhancement in chaperone activity of human $\alpha$A-crystallin by nanochaperone gold nanoparticles: Multispectroscopic studies on their molecular interactions', Spectrochim Acta A Mol Biomol Spectrosc, vol. 279, Oct. 2022, doi: 10.1016/J.SAA.2022.121344. Available: View Article
[18] M. Iosin, F. Toderas, P. L. Baldeck, and S. Astilean, 'Study of protein-gold nanoparticle conjugates by fluorescence and surface-enhanced Raman scattering', J Mol Struct, vol. 924-926, no. C, pp. 196-200, Apr. 2009, doi: 10.1016/J.MOLSTRUC.2009.02.004. Available: View Article
[19] H. Liu, N. Pierre-Pierre, and Q. Huo, 'Dynamic light scattering for gold nanorod size characterization and study of nanorod-protein interactions', Gold Bull, vol. 45, no. 4, pp. 187-195, Dec. 2012, doi: 10.1007/S13404-012-0067-4/FIGURES/5. Available: View Article

