Volume 12 - Year 2025 - Pages 79-87
DOI: 10.11159/jbeb.2025.010
Elucidation of Therapeutic Potential of Millet-Derived Peptides Inhibiting Fat Mass and Obesity-Associated Protein via In-Silico
Vinayak Kawale1, Sakshi Rai2#, Devraj J P2@, Ravindranath B S3, B Preethi4, Vankudavath Rajunaik2*
1School of Biotechnology, Amrita Vishwa Vidyapeetham, Kollam, Kerala State, India, vinayakkawale8060@gmail.com
2ICMR-National Institute of Nutrition, Tarnaka, Jamaiosmania PO, Hyderabad-500007, Telangana State, India
raisakshi722@gmail.com; jpdevraj26@gmail.com; *Corresponding Author: vivaswanraju@gmail.com
3Dept. Of Biotechnology, Manipal Institute of Technology, Manipal,
Manipal Academy of Higher Education, Manipal-576104, Udupi, Karnataka State, India
ravindranath.bs@manipal.edu
4Department of General Medicine, Bhaskar Medical College, Moinabad, Ranga Reddy-500075, Telangana State, India
9441236232preeti@gmail.com
#Sharing First Author, @Co-corresponding Author
Abstract - Over and undernutrition are generally perceived as lifestyle or diet related disorders. Apart from these external contributors, certain genes and proteins have been studied to plays a vital role in maintaining the metabolic state of an individual. One such gene is the Fat Mass and Obesity-Associated Protein (FTO) - an m6A RNA Demethylase, responsible for regulating energy homeostasis. This protein has been found to be strongly associated with obesity and related metabolic disorders. Targeting FTO with small-molecule inhibitors has shown promise as a therapeutic approach to manage obesity. The study employs a comprehensive computational strategy to identify bioactive peptides acting as potential natural inhibitors of the FTO protein derived from three millet species— Finger Millet (Eleusine Coracana), Pearl Millet (Pennisetum Glaucum), and Foxtail Millet (Setaria Italica). Bioactive peptides were curated from published literature focusing on millet seed proteins. Their physicochemical properties were assessed using PepCalc to evaluate stability and solubility. Subsequently, three-dimensional structures of the peptides were predicted using the I-TASSER server to generate high-confidence models for docking. Molecular docking analyses were conducted using ClusPro to examine Peptide-FTO binding affinities and interaction poses. The crystal structure of human FTO (retrieved from the Protein Data Bank) served as the docking target. Top-performing Peptide-FTO complexes were further taken for molecular dynamics (MD) simulations to evaluate the dynamic behavior and stability of the interactions. Key parameters such as RMSD, RMSF, and hydrogen bond profiles were analyzed over the course of the simulations. Our results show that several millet-derived peptides bind strongly and stably to the FTO protein, with favorable docking scores and sustained hydrogen bonding at its active site. These findings highlight the potential of millet peptides as natural FTO inhibitors for developing functional foods or nutraceuticals to combat obesity.
Keywords: FTO protein, millet-derived peptides, foxtail millet, finger millet, pearl millet, molecular docking, GROMACS, I-TASSER, ClusPro, PepCalc
© Copyright 2025 Authors This is an Open Access article published under the Creative Commons Attribution License terms. Unrestricted use, distribution, and reproduction in any medium are permitted, provided the original work is properly cited.
Date Received: 2025-04-10
Date Revised: 2025-09-30
Date Accepted: 2025-10-15
Date Published: 2025-11-25
1. Introduction
Cereal grains, including wheat, rice, barley, rye, oat, millet, and corn, have long been integral to the human diet. They contribute to a major portion of an individual’s daily intake, serving as primary sources of both macro- and micronutrients. Among cereals, millets stand out as nutrient-dense grains with a richer micronutrient profile enriched by bioactive flavonoids, fulfilling critical health demands. [1]. Millet proteins exclusively contain essential amino acids, majorly the sulphur-containing ones e.g., Methionine and Cysteine. Despite accounting for just 1.3% of total cereal production, millets are placed as the sixth most produced cereal. They are classified as minor, perennial grains of the Poaceae family, serving as staple foods in tropical and arid regions worldwide. Beyond their nutritional value, these archaeological staples are recognized for their medicinal properties derived from their content of vitamins, minerals, and bioactive compounds, earning them the designation of nutricereals. Millets can be categorized into major and minor types. In addition to exceptional nutritional attributes, multiple studies have demonstrated their therapeutic benefits, including reducing Prediabetes progression, improving Glycemic control, maintaining BMI, and lowering Cardiovascular risk [2],[3],[4].
Millets are Gluten-free and contain Resistant Starch (RS), while their Phytochemicals, such as Proanthocyanidins found in the grain and bran, have shown to possess anti-obesity effects by promoting satiety. Finger Millet (Eleusine Coracana), Pearl Millet (Pennisetum Glaucum), and Foxtail Millet (Setaria Italica) are nutrient-dense, ancient grains traditionally cultivated in arid and semi-arid regions of India and Africa. These grains are increasingly valued for their rich nutritional profiles, resilience to climate stress, and health-promoting properties. As modern diets face challenges from lifestyle disorders and nutrient deficiencies, Finger, Pearl, and Foxtail millets offer valuable functional foods that support health and sustainable food systems [5]. Finger millet contains Phenolic acids like Gallic, Ferulic, and P-coumaric acids, and Flavonoids such as Quercetin and Proanthocyanidins. Pearl millet is rich in Phenolic acids (147.8 mg/100g), Tocopherols, and Phytates, which in small amounts can also play protective roles. Other than compounds like Phytates, Flavonoids, and Tannins, many possible compounds can be of therapeutic interest to us. A supporting table mentioning major components of millet with roles in metabolic health has been provided in the Supplementary Table 1.
The BioActive compounds (BAs) in millets play crucial biochemical and physiological roles, significantly contributing to the management of metabolic disorders. These secondary metabolites affect various biological systems, providing numerous health benefits. Consumption of millets has been associated with reduced risks of Cardio Vascular Diseases (CVDs), Diabetes, and Hypertension. Specific compounds such as Luteolin, Taxifolin, Gallocatechin, Daidzein, Apigenin, and Procyanidins are implicated in Cardio Vascular protection [6]. Additionally, Violaxanthin, Zeaxanthin, Flavan-4-ol, and Daidzein are potent inhibitors of Human Angiotensin-Converting Enzyme (hACE). Phthalic acid, Naringenin, Daidzein, and Flavan-4-ol have been identified as potential inhibitors of Proprotein Convertase Subtilisin Kexin Type 9 (PCSK9) [7].
Obesity, characterized by chronic low-grade inflammation, is a major risk factor for metabolic syndrome and Insulin Resistance. An in vitro study reported Anti-Adipogenic effects of Polyphenol extracts from millets such as Proso, Little, and Pearl millet, which resulted in reduced intracellular lipid accumulation in differentiated 3T3-L1 cells. This effect involved the downregulation of transcription factors including PPAR, C/EBP, SREBP-1c, and pSREBP, alongside suppression of enzymes and regulators critical for fatty acid synthesis and metabolism, such as FAS, ACC, CDK2, ACOX1, and G6PD. These supporting evidence from the literature have built our understanding of millet bioactives and their role in metabolic regulation. Considering these as the fundamental basis, this study focuses on investigating millet-derived peptides as potential natural inhibitors of the fat mass and obesity-associated protein (FTO), a key regulator in Obesity pathways [8].
Fat Mass and Obesity-associated protein (FTO) is an m6A RNA Demethylase that has been implicated in the development of Obesity and Metabolic Disorders [9]. The protein influences multiple aspects of energy homeostasis and fat metabolism, making it a central player in the Pathogenesis of Obesity. Genetic variations in FTO affect appetite and satiety, while FTO's enzymatic activity impacts adipocyte function and energy expenditure, ultimately contributing to the development and maintenance of excess body weight and fat mass. Inhibition of FTO activity has emerged as a promising therapeutic strategy to manage Obesity. Bioactive peptides were curated from published literature focusing on millet seed proteins. Their Physicochemical properties were assessed using PepCalc to evaluate stability and solubility. Subsequently, three-dimensional structures of the peptides were predicted using the I-TASSER [10] server to generate high-confidence models for docking. Molecular docking analyses were conducted using ClusPro to examine peptide-FTO binding affinities and interaction poses. The crystal structure of human FTO (retrieved from the Protein Data Bank) served as the docking target. Top-performing peptide-FTO complexes were further subjected to Molecular Dynamics (MD) Simulations using GROMACS [11], [12] tool to evaluate the dynamic behavior and stability of the interactions. Key parameters such as RMSD, RMSF, and hydrogen bond profiles were analyzed over the course of the simulations. Our results show that several millet-derived peptides bind strongly and stably to the FTO protein, with favorable docking scores and sustained hydrogen bonding at its active site.
2. Material and Methods
2.1. Protein
The three-dimensional crystal structure of FTO protein from Homo Sapiens (PDB ID: 4IDZ) was obtained from the RCSB Protein Data Bank (PDB) (Fig 1). The selected structure consists of 495 amino acids in chain A and has a resolution of 2.46 Å, making it suitable for molecular interaction studies.
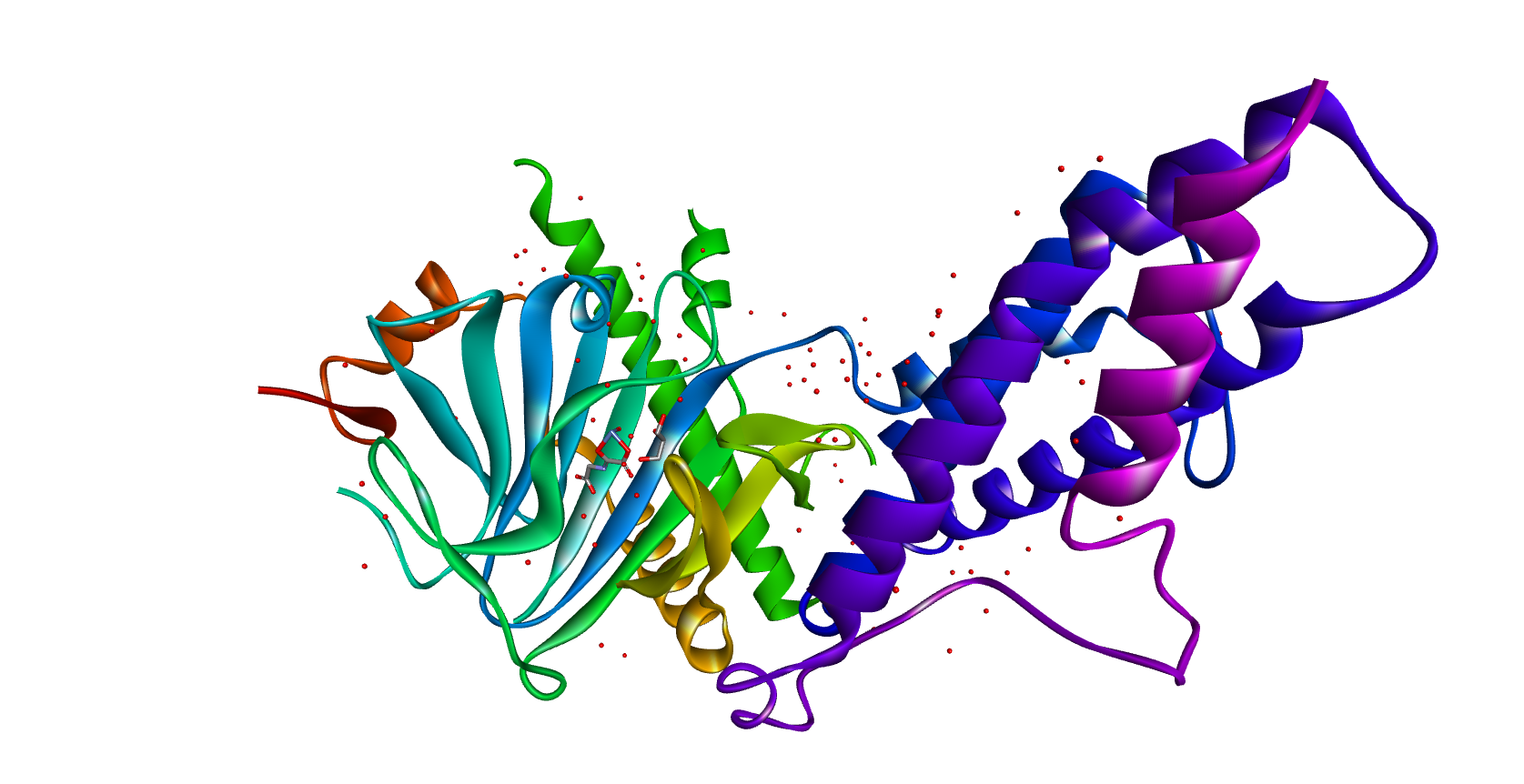
2.2. Peptide Collection and selection
Bioactive peptides were systematically collected through a literature-based approach, focusing on peer-reviewed research articles reporting peptide sequences derived from seed storage proteins of three millet species: Finger Millet (Eleusine Coracana), Pearl Millet (Pennisetum Glaucum), and Foxtail Millet (Setaria Italica)
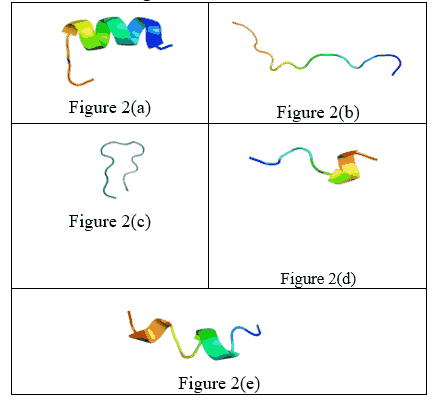
The selection was based on peptides with known or predicted bioactivities relevant to metabolic health. Firstly, from Finger Millet, two peptides, Fig 2(a) and Fig 2(b), as TSSSLNMAVRGGLTR and STTVGLGISMRSASVR were obtained [13]. These peptides were picked from a study exploring protein hydrolysates for their nutraceutical properties. Secondly, one peptide was selected from Pearl Millet, shown in Fig2 (c)- SDRDLLGPNNQYLPK. This was extracted from the work of Himani Agrawal, which characterized functional peptides derived from Pearl Millet proteins using enzymatic hydrolysis and mass spectrometry. Lastly, from Foxtail Millet, two peptides, shown as Fig 2(d) and Fig 2(e)- EDDQMDPMAK and QNWDFCEAWEPCF, respectively. These peptides were selected based on research by Hongbing Chen and Shuai Hu, which employed both in silico prediction and experimental validation [14].
These curated peptides formed the foundation for subsequent computational evaluations, including physicochemical characterization, structural modeling, molecular docking, and molecular dynamics simulations to assess their potential as inhibitors of the fat mass and obesity-associated (FTO) protein.
The selection criteria for peptide inclusion included parameters like- (i) Sequence length of sixteen amino acids or fewer to ensure computational tractability and potential oral bioavailability, (ii) Net charge between −3 and +2, allowing a balance between solubility and favorable electrostatic interactions with the FTO active site, (iii) Documented or predicted bioactivity (antioxidant, anti-obesity, or anti-inflammatory) from previous millet peptide studies, and (iv) Physicochemical stability as predicted by PepCalc. These criteria ensured a diverse set of peptide sequences with promising interaction potential and biological relevance.
2.3. Peptide Physicochemical Properties
The physicochemical properties of the collected peptides, including molecular weight (Mol. Wt.), net charge(Z), hydrophobicity, and isoelectric point (pI), were computed using the https://pepcalc.com/ web tool. These parameters provided initial insights into the stability, solubility, and potential bioactivity of the peptides.
Table 1. Peptide physiochemical properties
|
Sl. |
Number of residues |
Mol. Wt. (g/mol) |
pI |
Z at pH 7 |
Water solubility |
|
(a) |
Finger Millet Peptide-1 (15) |
1549.76 |
12.1 |
2 |
Good |
|
(b) |
Finger Millet Peptide-2 (16) |
1621.86 |
12.1 |
2 |
Poor |
|
(c) |
Pearl Millet Peptide-1 (15) |
1729.89 |
6.51 |
0 |
Good |
|
(d) |
Foxtail Millet Peptide-1 (10) |
1179.28 |
3.32 |
-3 |
Good |
|
(e) |
Foxtail Millet Peptide-2 (13) |
1674.81 |
0.65 |
-3.1 |
Poor |
2.4. Peptide Structure Prediction
The three-dimensional (3D) structures of the bioactive peptides collected from Finger Millet, Foxtail Millet, and Pearl Millet were predicted using the I-TASSER [10] webserver. The model with the highest C-score regardless of its rank was selected for downstream molecular docking and simulation studies.
2. 5. Molecular Docking
To investigate the interaction potential of millet-derived peptides with the fat mass and obesity-associated protein (FTO), molecular docking studies were performed using the ClusPro protein–protein docking server, a widely used tool for modeling peptide–protein interactions [15],[16]. Preparation of FTO protein is carried out using UCSF ChimeraX 1.10.1v [17]. The structure was refined by removing heteroatoms and water molecules, followed by the addition of hydrogen atoms to stabilize the protein for docking simulations. Any missing residues were identified and accounted for to ensure completeness and structural integrity of the target. Peptides were prepared by adding Hydrogen atoms to ensure accurate representation of potential bonding interactions during docking. Further, molecular docking was performed using the ClusPro 2.0v protein–protein docking server as it has been depicted in the adjacent figure. ClusPro performs rigid-body docking through Fast Fourier Transform (FFT)-based sampling and clusters them based on the lowest energy poses. The server evaluates docking poses using a weighted scoring function that approximates the binding energy. Docking results were ranked by cluster size and binding energy score, with the most populated and energetically favorable clusters selected for further analysis. The image shows the 3D interactions of the peptide with FTO. These peptides–FTO complexes were then subjected to molecular dynamics simulations to assess the stability and persistence of the interactions using LigPlot+2.3v [18].
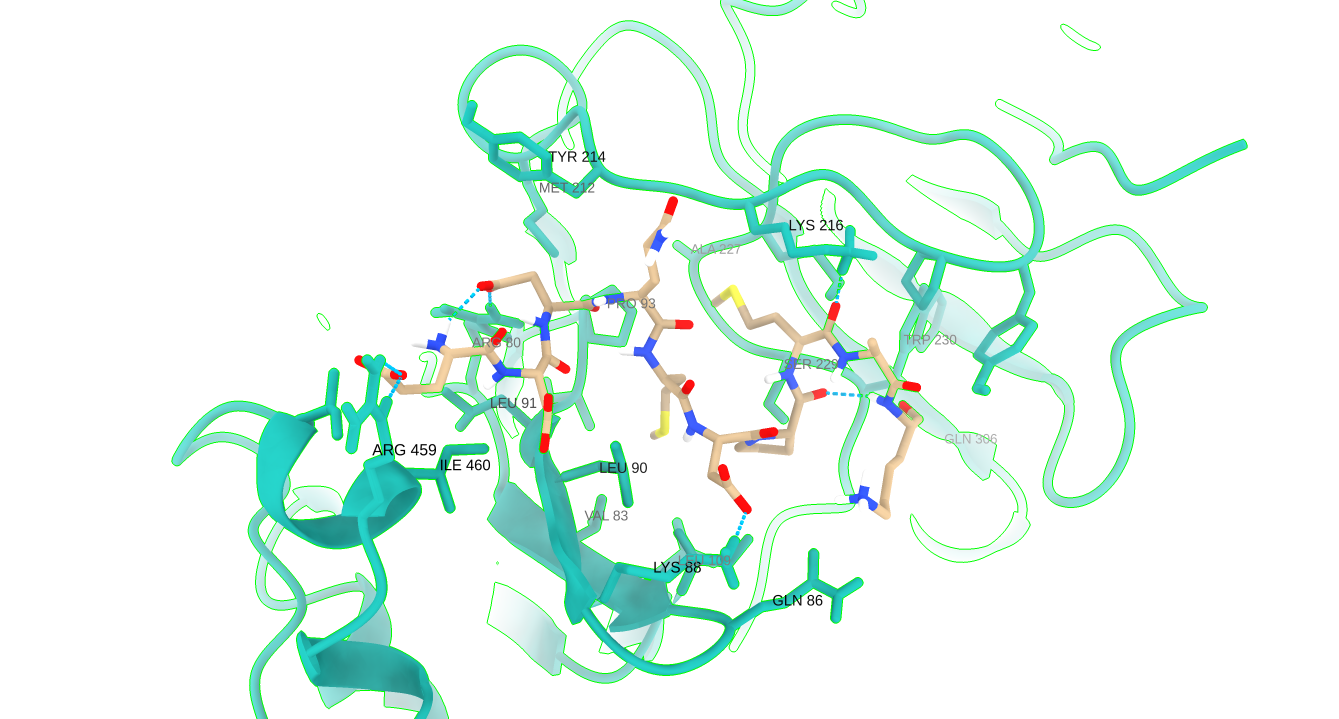
2. 6. Molecular Dynamics Simulation
To validate the docking results and assess the stability of the Peptide-FTO complexes, molecular dynamics simulations were conducted using GROMACS (GROningen MAchine for Chemical Simulations 2018v) [12], [11]. The simulation protocol involved solvating the systems, energy minimization, equilibration, and production runs to monitor complex behavior over time. To assess the conformational stability of the FTO–peptide complexes, Root Mean Square Deviation (RMSD) of the backbone atoms was calculated over a 100 ns molecular dynamics simulation. RMSD is a crucial parameter that provides insight into the extent of structural deviation from the initial conformation and serves as a key metric for evaluating system equilibration and structural integrity over time. Furthermore, root-mean-square fluctuation (RMSF), radius of gyration, hydrogen bond interactions, and binding free energies were analysed.
 |
|
|
|
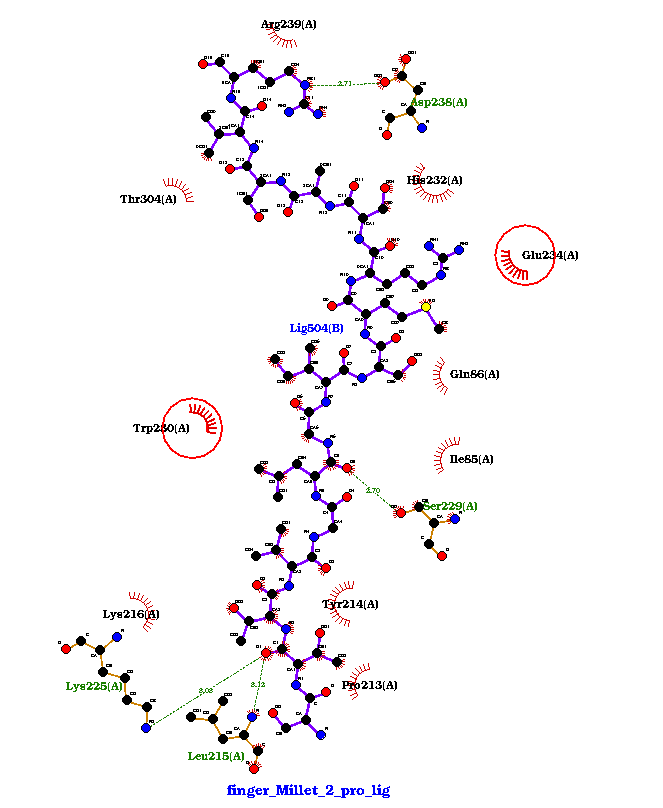
Fig 3 (a): Finger Millet 1: Binding Energy: -821.6, H-Bonds: 5 |
Fig 3 (b) Finger Millet 2: Binding Energy: -912.8, H-Bonds: 4 |
|
 |
|
|
|
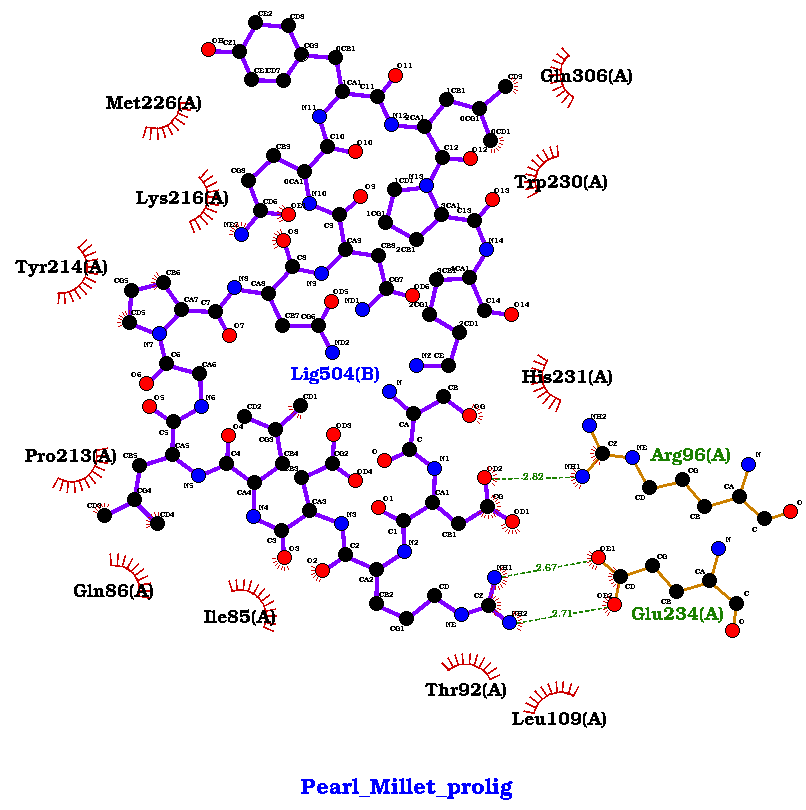
Fig 3 (c) Pearl Millet: Binding Energy : -655.8, H-Bonds: 3 |
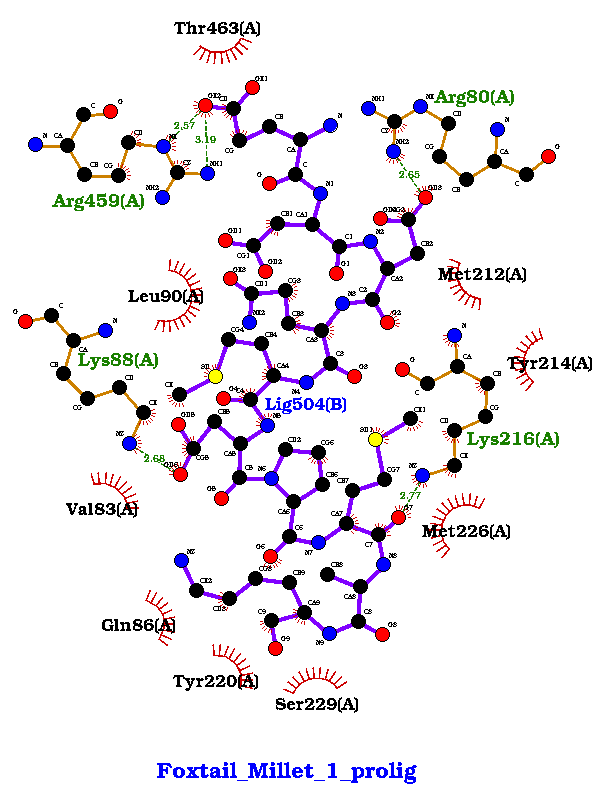
Fig 3 (d) Foxtail Millet-1: Binding Energy : -607, H-Bonds : 5 |
|
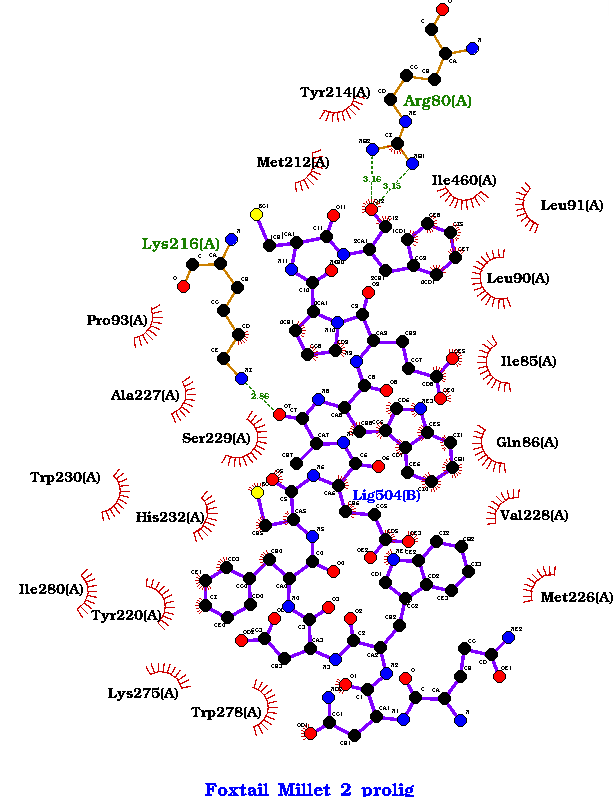 |
Fig 3 (e) Foxtail Millet-2: Binding Energy: -1133.4, H-Bonds: 3 |
|
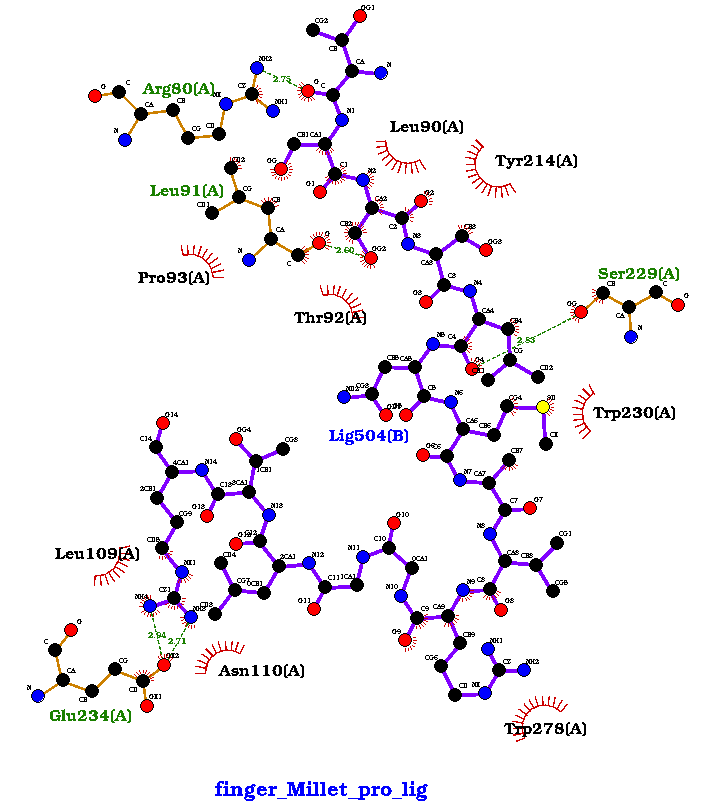
Figure 3. showing the 3D interactions of the peptide.
Fig 3 (a): Arg80, Leu90, Tyr214, Leu91, Tyr214, Pro93, Ther92, Ser229, Trp230, Trp278l, Asn110, Leu109.
Fig 3 (b): Arg239, Thr304, Asp238, His232, Glu234, Trp280, Gln86, Ile85, Ser229, Tyr214, Pro213, Leu215, Lys225, Lys216
Fig 3 (c): Met226, Gln306, Trp230, His231, Arg96, Glu234, Leu109, Thr92, Ile85, Gln86, Pro213, Tyr214, Lys216
Fig 3 (d): ArgA59, Thr463, Arg80, Met212, Tyr214, Lys216, Met226, Ser229, Tyr220, Gln86, Val83, Lys88, Leu90
Fig 3 (e): Tyr214, Met212, Ile460, Leu91, Leu90, Lys216, Ile85, Pro93, Ala227, Ser229, Gln86, Trp230, His232, Val228, Ile280, Tyr220, Met226, Lys275, Trp278
The molecular dynamics simulations were executed using the CHARMM36 Force Field within the GROMACS 2018.1 package. Each protein–peptide complex was placed in a Dodecahedral box maintaining a minimum distance of 1.0 nm between the solute and the box boundary and solvated using the TIP3P Water Model. Sodium and chloride ions were added to neutralize the system and achieve a physiological salt concentration of 0.15 M. Energy minimization was carried out using the Steepest Descent algorithm until the maximum force was below 1000 kJ/mol/nm. The system was equilibrated in two phases: a 100 ps NVT ensemble (constant volume and temperature) followed by a 100 ps NPT ensemble (constant pressure and temperature) using the V-rescale thermostat at 300 K and the Parrinello–Rahman barostat at 1 bar. A 100 ns production run was then conducted, and trajectories were saved every 2 ps for analysis. Periodic boundary conditions were applied in all directions, and the Particle Mesh Ewald (PME) method was used to calculate long-range electrostatic interactions.
3.Results
The computational analysis identified several peptides from Foxtail Millet, Finger Millet, and Pearl Millet with favorable binding affinity towards the FTO protein. The docking scores and simulation trajectories indicated stable interactions, with some peptides forming consistent hydrogen bonds with the active site residues of FTO. These findings suggest that millet-derived peptides have the potential to act as natural inhibitors of FTO, supporting their use in developing functional foods or therapeutics targeting obesity.
3.1 Molecular Dynamics Simulation Analysis of FTO-Peptide Complexes
3.1.1 RMSD Analysis
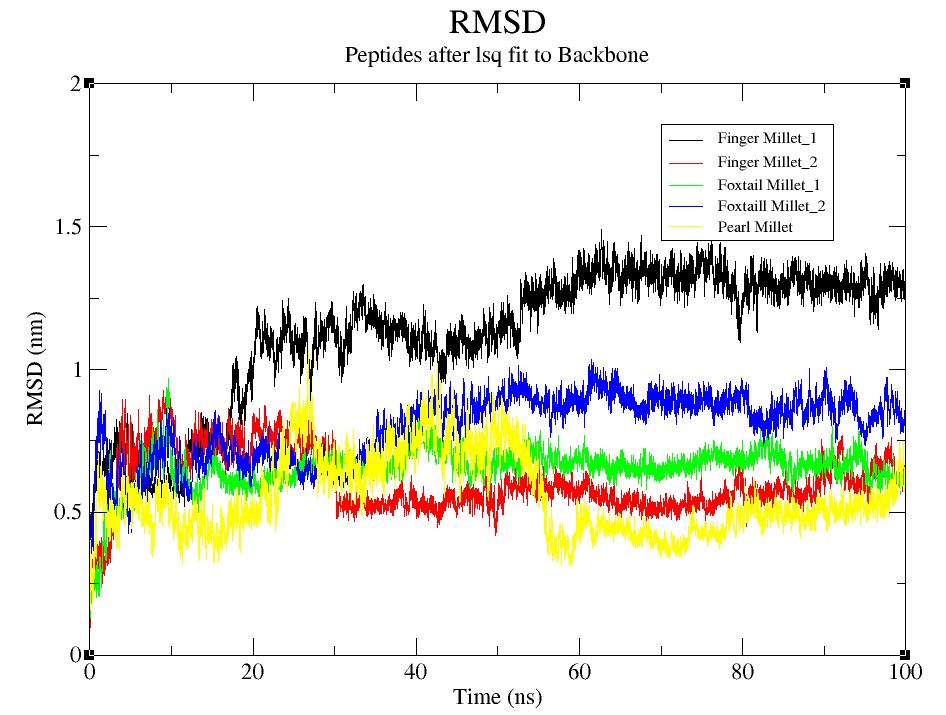
As illustrated in Fig 4(a), all peptide-bound FTO complexes underwent a characteristic RMSD stabilization phase within the first 20–30 ns, followed by consistent fluctuations, indicating that the systems reached equilibrium and remained dynamically stable. Among the complexes, Finger Millet_1 (black) exhibited the highest RMSD values (~1.4–1.5 nm), implying greater structural deviation and flexibility. Foxtail Millet_2 (blue) showed moderate RMSD levels (~0.8–1.0 nm), reflecting a stable yet relatively dynamic interaction. In contrast, Foxtail Millet_1 (green), Finger Millet_2 (red), and Pearl Millet (yellow) maintained lower RMSD values (~0.5–0.7 nm), denoting high conformational stability. Notably, the Pearl Millet complex remained consistently stable throughout the simulation, with minimal fluctuations, indicating strong peptide binding and low conformational drift. Similarly, Finger Millet_2 showed compact behavior with RMSD values remaining under 0.6 nm. These findings suggest that millet-derived peptides, particularly from Pearl and Foxtail Millet, form stable complexes with the FTO protein. The RMSD profiles highlight how sequence-specific interactions can influence structural dynamics and protein stability.
3.1.2 RMSF Analysis
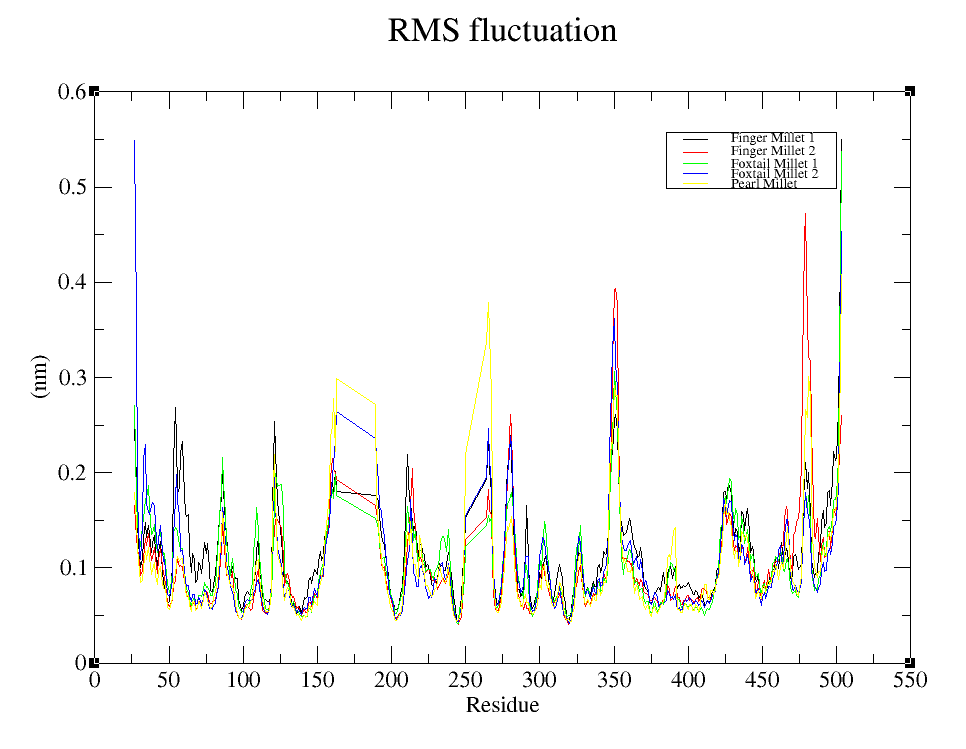
Root Mean Square Fluctuation (RMSF) analysis was conducted to evaluate the residue-wise flexibility of the FTO backbone in response to peptide binding. As shown in Fig 4(b), all complexes exhibited RMSF values between 0.1 and 0.35 nm for most residues, indicating a largely stable backbone conformation. Key fluctuations were observed at the N-terminal region (~residues 1–20), especially in Foxtail Millet_2 (blue), which exceeded 0.5 nm due to typical terminal flexibility. Minor peaks were also observed in loop regions around residues 180–190, 260–270, and 340–360, particularly in Pearl Millet (yellow) and Finger Millet_2 (red) complexes. Higher mobility was noted at the C-terminal end (~residues 500–520), with Finger Millet_2 showing the most pronounced fluctuation above 0.5 nm. The core region (residues 100–400) remained highly stable across all peptide-bound systems, with RMSF values under 0.2 nm. These results suggest that millet-derived peptides preserve the structural integrity of the FTO protein while permitting moderate flexibility in specific functional regions.
3.1.3 Radius of Gyration (Rg) Analysis
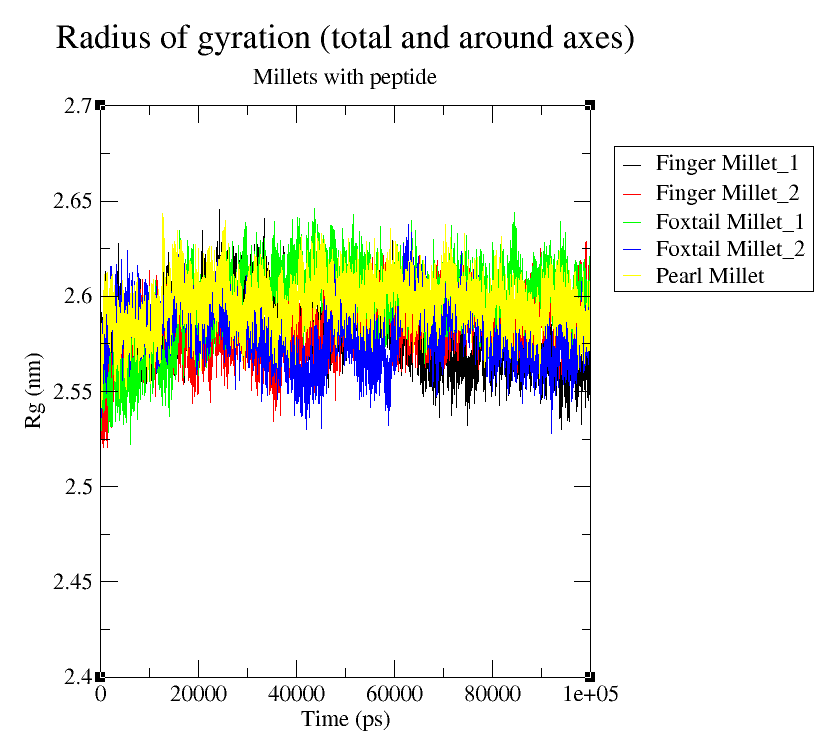
The radius of gyration (Rg) was calculated to evaluate the compactness and folding behavior of the peptide-bound FTO protein. As shown in Fig 4(c), all complexes maintained stable Rg values between 2.5 nm and 2.65 nm throughout the simulation. Pearl Millet (yellow) exhibited the highest average Rg (~2.63–2.66 nm), suggesting a slightly more extended protein conformation. Foxtail Millet_1 and Foxtail Millet_2 (green and blue) showed higher early-stage fluctuations but stabilized at ~2.6–2.63 nm. Finger Millet_1 and Finger Millet_2 (black and red) maintained lower Rg values (~2.52–2.57 nm), indicating more compact protein structures. These results demonstrate that peptide binding does not induce significant unfolding and that the structural compactness of FTO is maintained, reinforcing the stability of the modeled peptide–protein complexes.
3.1.4 Total Energy Analysis
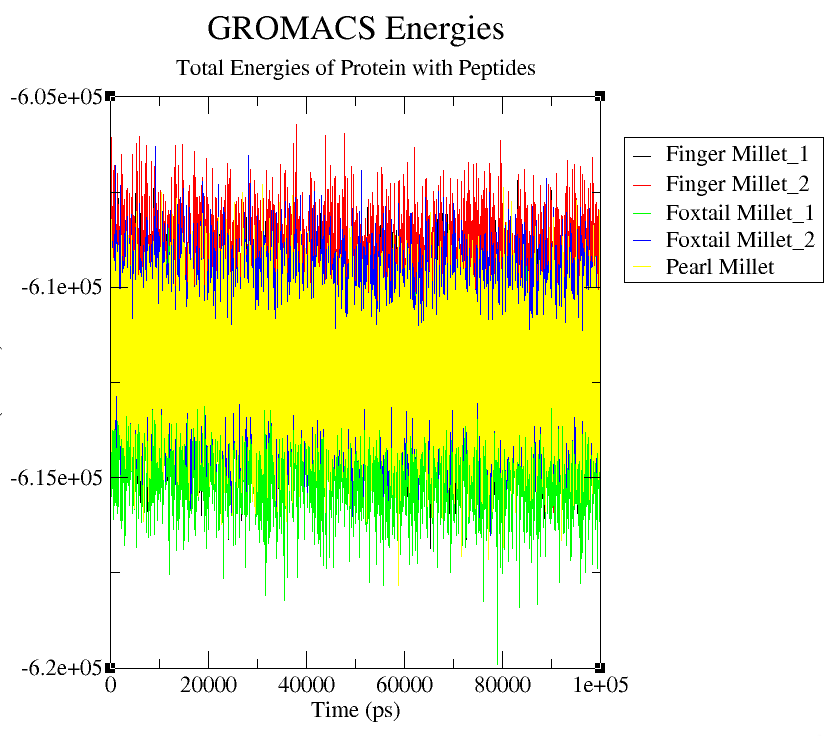
Total energy analysis was performed to examine the thermodynamic stability of the peptide–FTO complexes. As shown in Fig 4(d), all systems reached energetic equilibrium early in the simulation and remained within narrow energy ranges. Pearl Millet (yellow) displayed the most stable energy profile (−6.10 × 10⁵ kJ/mol), indicating strong and favourable peptide binding. Foxtail Millet peptides (green and blue) had slightly lower energies (−6.15 to −6.20 × 10⁵ kJ/mol), reflecting even more energetically stable interactions. Finger Millet peptides (black and red) showed slightly higher energies (~−6.05 to −6.10 × 10⁵ kJ/mol) but remained stable throughout. The consistency of energy values supports the structural and dynamic observations and suggests that millet-derived peptides contribute to the stability of FTO complexes, with foxtail and pearl millet peptides showing the most favourable binding profiles.
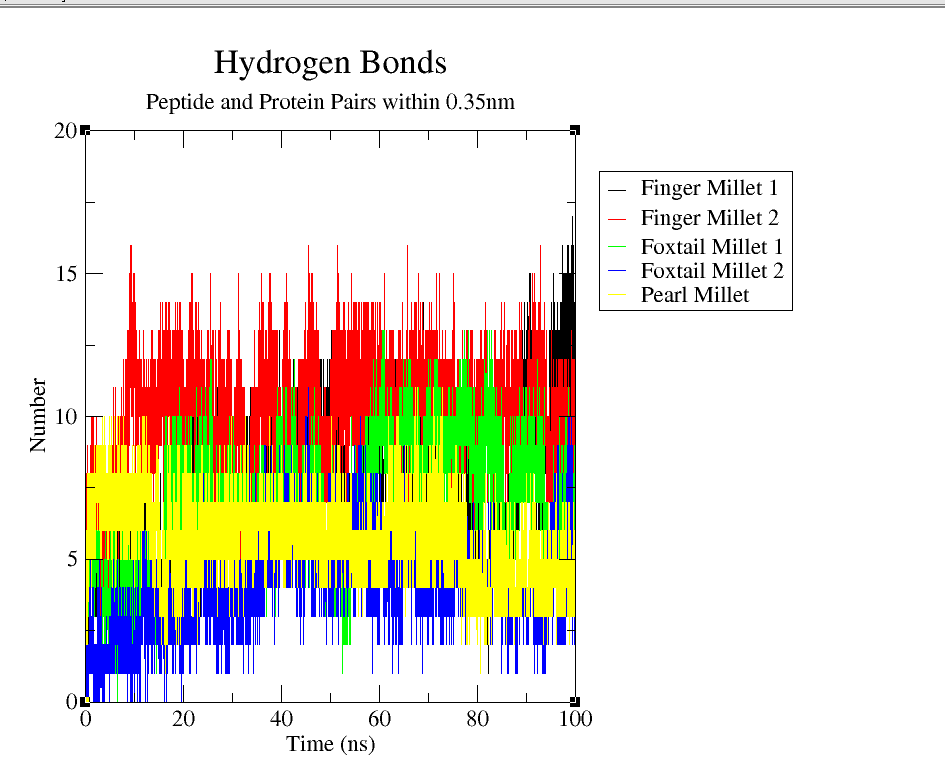
3.1.5 Hydrogen Bond Analysis
Hydrogen bonding interactions were assessed over the 100 ns simulation to quantify the binding affinity and interaction stability between FTO and the millet-derived peptides. A donor-acceptor distance cutoff of 0.35 nm was applied. As depicted in Fig 4(e), Finger Millet_2 (red) consistently formed the highest number of hydrogen bonds, averaging 10–15 and peaking near 18. Finger Millet_1 (black) also showed a strong interaction profile with 10–14 bonds. Foxtail Millet_1 (green) and Pearl Millet (yellow) formed moderate numbers of hydrogen bonds (6–10), while Foxtail Millet_2 (blue) exhibited the fewest (2–6), suggesting weaker or more transient interactions. These results highlight the variability in binding strength among the peptides and suggest that Finger Millet_2 and Pearl Millet are particularly effective in establishing stable hydrogen-bonded networks with the FTO protein.
4. Conclusion
This study utilized a comprehensive computational approach to identify millet-derived peptides as potential inhibitors of the FTO protein, a key player in obesity. Peptides from Finger, Foxtail, and Pearl Millet were modeled using I-TASSER and docked to FTO via ClusPro, revealing stable interactions. GROMACS-based molecular dynamics simulations confirmed the structural stability and binding affinity of the complexes. Pearl millet and finger millet 2 peptides showed the highest stability, with minimal RMSD and RMSF values, consistent Rg, and persistent hydrogen bonding. These findings highlight the therapeutic potential of millet peptides as natural FTO inhibitors, warranting further experimental validation.
Acknowledgement
We acknowledge the Director, Indian Council of Medical Research-National Institute of Nutrition, Hyderabad, Telangana State, India, for sponsoring and financially supporting the project.
References
[1] Seetha Anitha, Rosemary Botha, Joanna Kane-Potaka1, D Ian Givens, Ananthan Rajendran, Takuji W Tsusaka, Raj Kumar Bhandari, "Can Millet Consumption Help Manage Hyperlipidemia and Obesity?: A Systematic Review and Meta-Analysis," Front. Nutr., vol. 8, p. 700778, 2021, doi: 10.3389/fnut.2021.700778. Available: View Article
[2] Seetha Anitha, David Ian Givens, Rosemary Botha, Joanna Kane-Potaka,, Nur Liana Binti Sulaiman, Takuji W. Tsusaka, Kowsalya Subramaniam, Ananthan Rajendran, Devraj J. Parasannanavar, and Raj Kumar Bhandari "Calcium from Finger Millet-A Systematic Review and Meta-Analysis on Calcium Retention, Bone Resorption, and In Vitro Bioavailability." Accessed: Apr. 08, 2025. [Online]. Available: https://www.mdpi.com/2071-1050/13/16/8677 Available: View Article
[3] "IFCT_BOOK.pdf," Google Docs. Accessed: Apr. 08, 2025. [Online]. Available: https://drive.google.com/file/d/1tvpVuP3aaXs-6XIDr65GhE5RebWHq2gu/preview?usp=embed_facebook Available: View Document
[4] Jinu Jacob, Veda Krishnan, Chris Antony, Masimukka Bhavyasri, C Aruna, Kiran Mishra, Thirunavukkarasu Nepolean, Chellapilla Tara Satyavathi, Kurella B R S Visarada, "The nutrition and therapeutic potential of millets: an updated narrative review," Front. Nutr., vol. 11, p. 1346869, Apr. 2024, doi: 10.3389/fnut.2024.1346869. Available: View Article
[5] W. Zheng, C. Zhang, Y. Li, R. Pearce, E. W. Bell, and Y. Zhang, "Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations," Cell Rep. Methods, vol. 1, no. 3, p. 100014, Jul. 2021, doi: 10.1016/j.crmeth.2021.100014. Available: View Article
[6] Saini S, Saxena S, Samtiya M, Puniya M, Dhewa T. Potential of underutilized millets as Nutri-cereal: an overview. J Food Sci Technol. 2021 Dec;58(12):4465-77. Available: View Article
[7] Hithamani G, Ganesan P. Polyphenols from Indian cereal grains inhibit 3T3-L1 adipogenesis through modulating early and late phase adipogenic markers. Food Biosci. 2022 Dec 1;50:102075. Available: View Article
[8] Nagre K, Singh N, Meena A Probing the potential of bioactive compounds of millets as an inhibitor for lifestyle diseases: molecular docking and simulation-based approach. Available: View Article
[9] H. Agrawal, R. Joshi, and M. Gupta, "Isolation, purification and characterization of antioxidative peptide of pearl millet (Pennisetum glaucum) protein hydrolysate," Food Chem., vol. 204, pp. 365-372, Aug. 2016, doi: 10.1016/j.foodchem.2016.02.127. Available: View Article
[10] S. Hu and J. Yuan, "Antioxidant and Anti-Inflammatory Potential of Peptides Derived from In Vitro Gastrointestinal Digestion of Germinated and Heat-Treated Foxtail Millet (Setaria italica) Proteins | Journal of Agricultural and Food Chemistry." Accessed: Apr. 08, 2025. [Online]. Available: https://pubs.acs.org/doi/10.1021/acs.jafc.0c03732 Available: View Article
[11] George Jones, Akhil Jindal, Usman Ghani, Sergei Kotelnikov, Megan Egbert, Nasser Hashemi, Sandor Vajda, Dzmitry Padhorny, Dima Kozakov, "Elucidation of protein function using computational docking and hotspot analysis by ClusPro and FTMap," Acta Crystallogr. Sect. Struct. Biol., vol. 78, no. 6, pp. 690-697, Jun. 2022, doi: 10.1107/S2059798322002741. Available: View Article
[12] D. Kozakov, D. Hall, and B. Xia, "The ClusPro web server for protein-protein docking | Nature Protocols." Accessed: Apr. 08, 2025. [Online]. Available: https://www.nature.com/articles/nprot.2016.169 Available: View Article
[13] Eric F Pettersen, Thomas D Goddard, Conrad C Huang, Elaine C Meng, Gregory S Couch, Tristan I Croll, John H Morris, Thomas E Ferrin, "UCSF ChimeraX: Structure visualization for researchers, educators, and developers," Protein Sci. Publ. Protein Soc., vol. 30, no. 1, pp. 70-82, Jan. 2021, doi: 10.1002/pro.3943. Available: View Article
[14] R. A. Laskowski and M. B. Swindells, "LigPlot+: multiple ligand-protein interaction diagrams for drug discovery," J. Chem. Inf. Model., vol. 51, no. 10, pp. 2778-2786, Oct. 2011, doi: 10.1021/ci200227u Available: View Article

